Genome assembly2
From CSBLwiki
Contents |
Lactobacillus genus
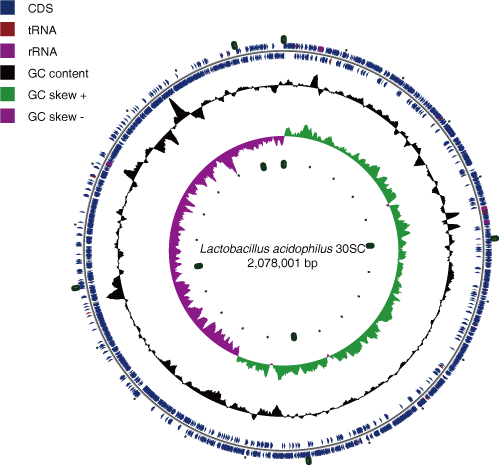
Circular view
copy # and position of rRNA operon
- copy # : 4
- position
- 2h,3h, 5t, 7h, 9h, 11h, 13h
Terminus
- scf13 : oriC, 155nt, 56722..56876 nt
- scf10_4
rRNA operon of Other species
| Name | rRNA operon | Total Len | transposase |
| Lactobacillus_acidophilus_NCFM | 4 | 2M | 40 |
| Lactobacillus_brevis_ATCC_367 | 5 | 2.2M | |
| Lactobacillus_casei | 5 | 3M | |
| Lactobacillus_casei_ATCC_334 | 5 | 2.9M | |
| Lactobacillus_casei_Zhang_uid50673 | 5 | 2.8M | |
| Lactobacillus_crispatus_ST1_uid48359 | 4 | 2M | |
| Lactobacillus_delbrueckii_bulgaricus | 9 | 1.8M | |
| Lactobacillus_delbrueckii_bulgaricus_ATCC_BAA-365 | 9 | 1.8M | |
| Lactobacillus_fermentum_IFO_3956 | 5 | 2.1M | |
| Lactobacillus_gasseri_ATCC_33323 | 6 | 1.9M | |
| Lactobacillus_helveticus_DPC_4571 | 4 | 2.1M | 260 |
| Lactobacillus_johnsonii_FI9785 | 4 | 1.8M | |
| Lactobacillus_johnsonii_NCC_533 | 6 | 2M | |
| Lactobacillus_plantarum | 5 | 3.3M | |
| Lactobacillus_plantarum_JDM1 | 5 | 3.2M | |
| Lactobacillus_reuteri_DSM_20016 | 6 | 2M | |
| Lactobacillus_reuteri_F275_Kitasato | 6 | 2M | |
| Lactobacillus_rhamnosus_GG | 5 | 3M | |
| Lactobacillus_rhamnosus_Lc_705 | 5 | 3M | |
| Lactobacillus_sakei_23K | 7 | 1.9M | |
| Lactobacillus_salivarius_UCC118 | 7 | 1.8M |
Read
| Flatform | Read Type | Total Reads | Number of Reads Used | Number of Bases Used | Percent Reads Assembled | Percent Bases Assembled |
| Solexa Illumina | SE | 5359073 | - | - | - | - |
| Fake Reads(FR) (Solexa/Illumina) | SE | 8390 | 11964248 | 8243 | 98.25 | 95.15 |
| FR (CABOG) | SE | 7020 | 10529913 | 7017 | 99.96 | 99.93 |
| Roche 454 | PE | 158188 (270784) | ||||
| Roche 454 | PE | 235924 (364291) |
- Solexa and 454 derived from different strain.
- Only use 454 reads.
Assembly flow
- illumina -> ABySS: 4336 contigs, total 4M; very bad -> fake reads
- fake reads(ABySS illumina) + 454reads -> Newbler : 443 scaffolds; very bad => reject
- only 454 -> Newbler,gapResolution -> 15scf, 21ctg -> Running PCR NOW
- only 454 -> CABOG -> compare(mapping) to Newbler (by nucmer & mummerplot) -> some disagree in scaffolds between 2 softwares
- CABOG can assemble rRNA operon
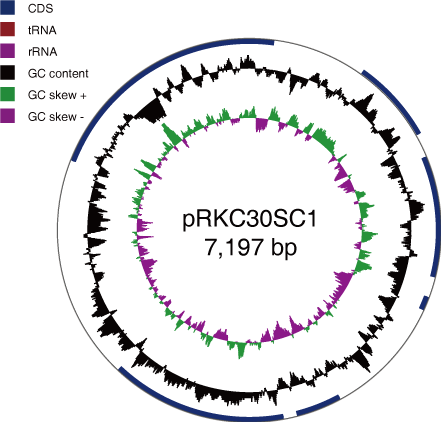
- found plasmid
- gap 10_2-10_3: filled by CABOG contig
Assembly Result
| Assembler | Contig Type | Number of Contigs | Total bases | |
| velvet | contigs | 14702 | 2674691 | |
| ABySS | Large Contig (>500bp) | 4336 | 4121677 | *very bad |
| Newbler(FR(Sol/Ill) + 454 | Scaffolds | 443 | 4585959 | *very bad, reject |
| Newbler(454 only), gapResolution | Scaffolds | 15 | 2058137 | Running PCR |
| Newbler(454 only), gapResolution | Contigs | 21 | 2053877 | Running PCR |
| CABOG(454 only) | Scaffolds (>500bp) | 13 | 2120461 | |
| CABOG(454 only) | Contigs (>500bp) | 26 | 2119042 | |
| Newbler(FR(CABOG) +454) | Scaffolds | 7 | 2051269 | |
| Newbler(FR(CABOG) +454) | Scaffolds | 59 | 2012620 |
- 1 chromosome and 2 plasmids are assembled
- 1986681bp, 7197bp, 12568bp
Links
Scripts
Etc
- NCBI Genomes
- Lactobacillus delbrueckii bulgaricus by Genoscope
- Tech Summary: Illumina's Solexa Sequencing Technology
- Case stuides:
- Gene-Boosted Assembly of a Novel Bacterial Genome from Very Short Reads
- High-Precision, Whole-Genome Sequencing of Laboratory Strains Facilitates Genetic Studies
- De novo bacterial genome sequencing: Millions of very short reads ...
- Short read fragment assembly of bacterial genomes...
- De novo fragment assembly with short mate-paired reads: Does the read length matter? (PDF)
- Solexa format & Fastq format
- Benchmark papers
- A Draft Genome Sequence of Pseudomonas syringae pv. tomato T1 Reveals a Type III Effector Repertoire Significantly Divergent from That of Pseudomonas syringae pv. tomato DC3000, MPMI(2008) (PDF)
- High-throughput sequencing provides insights into genome variation and evolution in Salmonella Typhi, Nature Genetics (2008)