ComGen Course
From CSBLwiki
(Difference between revisions)
(→To check=) |
(→Languages) |
||
| Line 76: | Line 76: | ||
*Biopython ([http://biopython.org/wiki/Biopython Download]) | *Biopython ([http://biopython.org/wiki/Biopython Download]) | ||
**[http://biopython.org/DIST/docs/tutorial/Tutorial.html Tutorial](follow the instruction) | **[http://biopython.org/DIST/docs/tutorial/Tutorial.html Tutorial](follow the instruction) | ||
| + | *[http://matplotlib.sourceforge.net/users/pyplot_tutorial.html Pyplot Tutorial (matplotlib)] | ||
| + | *[http://www.scipy.org/Tentative_NumPy_Tutorial NumPy Tutorial] | ||
| + | |||
===Packages=== | ===Packages=== | ||
*R ([http://cran.r-project.org R packages]) | *R ([http://cran.r-project.org R packages]) | ||
**[http://cran.r-project.org/manuals.html Manuals] | **[http://cran.r-project.org/manuals.html Manuals] | ||
**[http://cran.r-project.org/other-docs.html Contributed Documents] | **[http://cran.r-project.org/other-docs.html Contributed Documents] | ||
Revision as of 15:21, 19 March 2009
Contents |
Class schedule
Chapter Assign Pages Presentation Due date 1 이은혜 21 03/19/09 03/12/09 2 박애경 16 03/26/09 03/21/09 3 고혁진 23 04/02/09 03/26/09 4 장은혁 17 04/07/09 04/02/09 5 이예림 18 04/16/09 04/07/09 6 김소현 14 04/23/09 04/16/09 7 정진아 18 05/14/09 04/23/09 8 김윤식 12 05/21/09 04/30/09 9 김윤식 18 06/04/09 05/07/09 10 김윤식 21 06/11/09 05/14/09
- No Class
- 4/30 (중간고사)
- 5/7 (학회참석, SF)
- 5/28 (학회참석, Cheju)
- 해당 단원은 발표 1주일전에 EKU에 올려 놓을것 (MS-Word 형식으로 제출)
- 발표는 해당 단원의 소개 및 요약
- 각 단원의 연습문제를 풀어서 제출할것 - EKU
- 발표한 내용을 MS-Word의 "Trace Changes" 기능을 이용하여 수정하여 제출
Chapters
Chapter 1
- Reading (download a PDF) by 이은혜
- Installing Python & related Modules (Windows & Linux only)
- Python(x,y)-2.1.11 - Free scientific and engineering development software download & install (current version is 2.1.1; 3/19/2009) - including very useful scientific modules (Numpy, Scipy...)
- Download from local deposit (Python(x,y)-2.1.11.exe)
- Biopython 1.49 download biopython-1.49.win32-py2.5.exe
Excercise#1 Download a genome sequence & do basic statistical analysis
- GC-content?
- ANS: GC content of NC_01415 is '49.9%'
- Code
>>> from Bio import Entrez, SeqIO
>>> handle = Entrez.efetch(db="nucleotide",id="NC_001416",rettype="fasta")
>>> record = SeqIO.read(handle,"fasta")
>>> print record
ID: gi|9626243|ref|NC_001416.1|
Name: gi|9626243|ref|NC_001416.1|
Description: gi|9626243|ref|NC_001416.1| Enterobacteria phage lambda, complete genome
Number of features: 0
Seq('GGGCGGCGACCTCGCGGGTTTTCGCTATTTATGAAAATTTTCCGGTTTAAGGCG...ACG', SingleLetterAlphabet())
>>> print len(record)
48502
>>> record
SeqRecord(seq=Seq('GGGCGGCGACCTCGCGGGTTTTCGCTATTTATGAAAATTTTCCGGTTTAAGGCG...ACG', SingleLetterAlphabet()), id='gi|9626243|ref|NC_001416.1|', name='gi|9626243|ref|NC_001416.1|', description='gi|9626243|ref|NC_001416.1| Enterobacteria phage lambda, complete genome', dbxrefs=[])
>>> record.seq
Seq('GGGCGGCGACCTCGCGGGTTTTCGCTATTTATGAAAATTTTCCGGTTTAAGGCG...ACG', SingleLetterAlphabet())
>>> from Bio.SeqUtils import GC
>>> GC(record.seq)
49.857737825244321
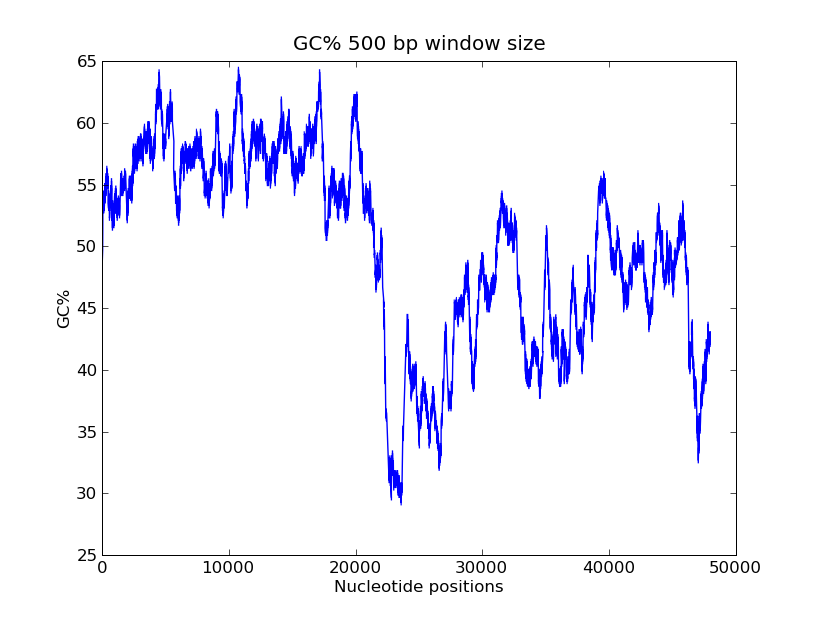
- GC-content scanning with window size 500 bps?
- ANS:
- Code
>>>x = record.seq
>>>windowsize = 500
>>>gc_values = [ GC(x[i:(i+499)] for i in range(1,len(x)-windowsize+1) ]
>>> import pylab
>>> pylab.plot(gc_values)
>>> pylab.title("GC% 500 bp window size")
>>> pylab.xlabel("Nucleotide positions")
>>> pylab.ylabel("GC%")
>>> pylab.show()
A Thinking Chair
- independent and identically distributed (i.i.d.)
Programming
Languages
- Python (Official Site)
- Biopython (Download)
- Tutorial(follow the instruction)
- Pyplot Tutorial (matplotlib)
- NumPy Tutorial