Synbio course
From CSBLwiki
(Difference between revisions)
(→Practice #1) |
(→Practice #2) |
||
| (29 intermediate revisions not shown) | |||
| Line 1: | Line 1: | ||
| + | {| align=right cellpadding=15 | ||
| + | |__TOC__ | ||
| + | |} | ||
==2010 Fall LMB904== | ==2010 Fall LMB904== | ||
| - | *[[Synthetic_Biology|Introduction to Synthetic Biology]] | + | *What is Synthetic Biology? - [[Synthetic_Biology|Introduction to Synthetic Biology]] |
| - | *[[R|R package]] | + | **check readings (recent special issue of various journals) in above link |
| - | + | ||
| - | **tutorials and course materials in [[R|R page]] | + | *We will use [[R|R package]] for simulating several biological processes |
| - | *We will practice | + | **check [http://bioconductor.org bioconductor] for more applications in bioinformatics |
| + | **tutorials and course materials in CSBL's [[R|R page]] (some collections) | ||
| + | |||
| + | ==Practices in R== | ||
| + | *We will practice some materials in [http://ocw.mit.edu/courses/physics/8-591j-systems-biology-fall-2004/readings/ MIT open courseware] (department of physics) | ||
===Practice #1=== | ===Practice #1=== | ||
| - | *Simulating Michaelis-Menten kinetics in R | + | *Simulating [[w:Michaelis-Menten kinetics|Michaelis–Menten kinetics]] in R |
| + | **underlying assumption: quasi-equilibrium state (pseudosteady state) of enzyme-catalyzed reaction | ||
| + | **[http://ocw.mit.edu/courses/physics/8-591j-systems-biology-fall-2004/readings/CodeI1_v2.m OCW Matlab code] will be ported in R | ||
| + | **using R | ||
| + | ***install a specific package (e.g. 'odesolve') in R (there are several ways to do this) | ||
| + | <pre> | ||
| + | # execute 'R' & run a command | ||
| + | # you should have an internet connection | ||
| + | |||
| + | >install.package("odesolve") | ||
| + | |||
| + | </pre> | ||
| + | |||
| + | *R-code | ||
| + | <pre> | ||
| + | library(odesolve) | ||
| + | # Michaelis-menten equation | ||
| + | # E + S <-> ES -> E + P | ||
| + | # k1,k2 k3 | ||
| + | # rate equations | ||
| + | # dE = -k1*E*S+k2*ES, dS = -k1*E*S+k2*ES+k3*ES, dES = k1*E*S-k2*ES-k3*ES | ||
| + | # quasi-equilibrium; E0 = E + ES; dES = 0 | ||
| + | # dE = -k1*E0*S+(k1*S+k2)*ES, dES = k1*E0*S-(k1*S+k2+k3)*ES, dP = k2*ES | ||
| + | |||
| + | ### define ode functions | ||
| + | michaelis = function(t,y,p) { | ||
| + | # t, y, p: vector | ||
| + | # t: a time scale | ||
| + | # y: S = y[1]; ES = y[2]; P = y[3] | ||
| + | # p: k1 = p[1]; k2 = p[2]; k3=p[3]; E0=p[4] | ||
| + | dS = -p[1]*p[4]*y[1]+(p[1]*y[1]+p[2])*y[2] | ||
| + | dES = p[1]*p[4]*y[1]-(p[1]*y[1]+p[2]+p[3])*y[2] | ||
| + | dP = p[2]*y[2] | ||
| + | list(c(dS,dES,dP)) | ||
| + | } | ||
| + | |||
| + | #### initial parameters | ||
| + | k1 = 1e3 | ||
| + | k2 = 1 | ||
| + | k3 = 5e-2 | ||
| + | E0 = 5e-4 | ||
| + | |||
| + | p = c(k1,k2,k3,E0) # <-- parameters (initial values) | ||
| + | t = seq(0,100,1) # <-- a time scale (0 to 100 sec.) | ||
| + | |||
| + | y = c(1e-3,0,0) # <-- initial values of [S], [ES] and [P] | ||
| + | |||
| + | #### solving equations | ||
| + | res = lsoda(y,t,michaelis,p) # store result (101 x 4 matrix) | ||
| + | |||
| + | |||
| + | #### plotting results | ||
| + | time = res[,1] # time range - column 1 | ||
| + | S = res[,2] # [S] changes - column 2 | ||
| + | ES = res[,3] # [ES] | ||
| + | E = p[4]-res[,3] # [E] free enzyme | ||
| + | P = res[,4] # [P] product | ||
| + | |||
| + | plot(time,S,type="l",col="red",ylim=c(0,0.001)) | ||
| + | points(time,ES,type="l",col="blue") | ||
| + | points(time,E,type="l",col="cyan") | ||
| + | points(time,P,type="l",col="green") | ||
| + | </pre> | ||
| + | |||
| + | *Result (above script is working but the result will not be correct; you may have to find an error in the code) | ||
| + | {| align=center | ||
| + | |- | ||
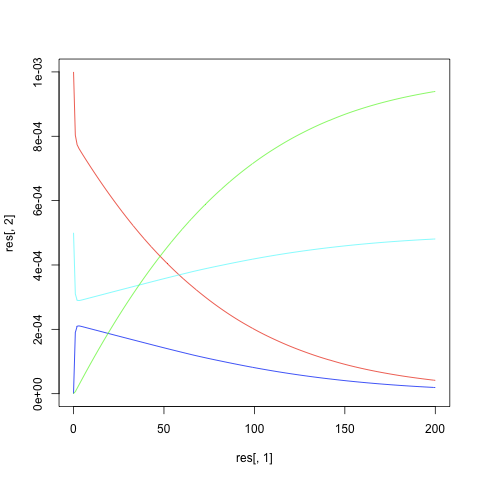
| + | |[[image:Michaelis-menten.png|thumb|left|Michaelis-Menten simulation result (correct one)]] | ||
| + | |} | ||
| + | |||
===Practice #2=== | ===Practice #2=== | ||
*A Genetic Switch in Lamba Phage in R | *A Genetic Switch in Lamba Phage in R | ||
| + | <pre> | ||
| + | library(odesolve) | ||
| + | # Lambda phage lysogeny | ||
| + | # ODE | ||
| + | # dx/dt = alpha*x^2/(1+(1+sigma1)*x^2+sigma2*x^4)-gamma*x+1 | ||
| + | # | ||
| + | # define ODE function | ||
| + | hastyfunc = function(t,y,p) { | ||
| + | # p[1:4] = c(alpha,gamma,sigma1,sigma2) | ||
| + | dy = p[1]*y^2/(1+(1+p[3])*y^2+p[4]*y^4)-p[2]*y+1 | ||
| + | list(dy) | ||
| + | } | ||
| + | # | ||
| + | # initial parameters | ||
| + | # | ||
| + | # p[1:4] = c(alpha,gamma,sigma1,sigma2) | ||
| + | p = c(50,20,1,5) | ||
| + | t = seq(0,10,1) | ||
| + | y0 = 0; y1 = 1 | ||
| + | res0 = lsoda(y0,t,hastyfunc,p) | ||
| + | res1 = lsoda(y1,t,hastyfunc,p) | ||
| + | plot(t,res0[,2],type="l",col="blue",main="Hasty function", | ||
| + | xlab="Time",ylim=c(0,1)) | ||
| + | points(t,res1[,2],type="l",col="red") | ||
| + | </pre> | ||
<biblio>LAM pmid=10681449</biblio> | <biblio>LAM pmid=10681449</biblio> | ||
| + | |||
===Practice #3=== | ===Practice #3=== | ||
*A Genetic Toggle Switch in R | *A Genetic Toggle Switch in R | ||
<biblio>TOG pmid=10659857</biblio> | <biblio>TOG pmid=10659857</biblio> | ||
| + | <!-- <pubmed>10659857</pubmed> --> | ||
Latest revision as of 14:47, 18 November 2010
|
2010 Fall LMB904
- What is Synthetic Biology? - Introduction to Synthetic Biology
- check readings (recent special issue of various journals) in above link
- We will use R package for simulating several biological processes
- check bioconductor for more applications in bioinformatics
- tutorials and course materials in CSBL's R page (some collections)
Practices in R
- We will practice some materials in MIT open courseware (department of physics)
Practice #1
- Simulating Michaelis–Menten kinetics in R
- underlying assumption: quasi-equilibrium state (pseudosteady state) of enzyme-catalyzed reaction
- OCW Matlab code will be ported in R
- using R
- install a specific package (e.g. 'odesolve') in R (there are several ways to do this)
# execute 'R' & run a command
# you should have an internet connection
>install.package("odesolve")
- R-code
library(odesolve)
# Michaelis-menten equation
# E + S <-> ES -> E + P
# k1,k2 k3
# rate equations
# dE = -k1*E*S+k2*ES, dS = -k1*E*S+k2*ES+k3*ES, dES = k1*E*S-k2*ES-k3*ES
# quasi-equilibrium; E0 = E + ES; dES = 0
# dE = -k1*E0*S+(k1*S+k2)*ES, dES = k1*E0*S-(k1*S+k2+k3)*ES, dP = k2*ES
### define ode functions
michaelis = function(t,y,p) {
# t, y, p: vector
# t: a time scale
# y: S = y[1]; ES = y[2]; P = y[3]
# p: k1 = p[1]; k2 = p[2]; k3=p[3]; E0=p[4]
dS = -p[1]*p[4]*y[1]+(p[1]*y[1]+p[2])*y[2]
dES = p[1]*p[4]*y[1]-(p[1]*y[1]+p[2]+p[3])*y[2]
dP = p[2]*y[2]
list(c(dS,dES,dP))
}
#### initial parameters
k1 = 1e3
k2 = 1
k3 = 5e-2
E0 = 5e-4
p = c(k1,k2,k3,E0) # <-- parameters (initial values)
t = seq(0,100,1) # <-- a time scale (0 to 100 sec.)
y = c(1e-3,0,0) # <-- initial values of [S], [ES] and [P]
#### solving equations
res = lsoda(y,t,michaelis,p) # store result (101 x 4 matrix)
#### plotting results
time = res[,1] # time range - column 1
S = res[,2] # [S] changes - column 2
ES = res[,3] # [ES]
E = p[4]-res[,3] # [E] free enzyme
P = res[,4] # [P] product
plot(time,S,type="l",col="red",ylim=c(0,0.001))
points(time,ES,type="l",col="blue")
points(time,E,type="l",col="cyan")
points(time,P,type="l",col="green")
- Result (above script is working but the result will not be correct; you may have to find an error in the code)
Practice #2
- A Genetic Switch in Lamba Phage in R
library(odesolve)
# Lambda phage lysogeny
# ODE
# dx/dt = alpha*x^2/(1+(1+sigma1)*x^2+sigma2*x^4)-gamma*x+1
#
# define ODE function
hastyfunc = function(t,y,p) {
# p[1:4] = c(alpha,gamma,sigma1,sigma2)
dy = p[1]*y^2/(1+(1+p[3])*y^2+p[4]*y^4)-p[2]*y+1
list(dy)
}
#
# initial parameters
#
# p[1:4] = c(alpha,gamma,sigma1,sigma2)
p = c(50,20,1,5)
t = seq(0,10,1)
y0 = 0; y1 = 1
res0 = lsoda(y0,t,hastyfunc,p)
res1 = lsoda(y1,t,hastyfunc,p)
plot(t,res0[,2],type="l",col="blue",main="Hasty function",
xlab="Time",ylim=c(0,1))
points(t,res1[,2],type="l",col="red")
Error fetching PMID 10681449:
- Error fetching PMID 10681449:
Practice #3
- A Genetic Toggle Switch in R
Error fetching PMID 10659857:
- Error fetching PMID 10659857: