E. coli knockout
From CSBLwiki
(Difference between revisions)
(→Procedure) |
(→Procedure) |
||
| Line 41: | Line 41: | ||
***Sequences from cassette region have about 60℃ Tm-value. | ***Sequences from cassette region have about 60℃ Tm-value. | ||
***Overview is attached. [[media:overview.jpg|File:Overview]] | ***Overview is attached. [[media:overview.jpg|File:Overview]] | ||
| - | *PCR condition | + | **PCR condition |
| - | ** Anneling temparature and extension time is depend on the target sequence. | + | *** Anneling temparature and extension time is depend on the target sequence. |
| - | ** Anneling temparature is tested at 55℃ at first. | + | *** Anneling temparature is tested at 55℃ at first. |
| - | ** Extension time is depend on the length of the target sequence. (1kb = about 1 min.) | + | *** Extension time is depend on the length of the target sequence. (1kb = about 1 min.) |
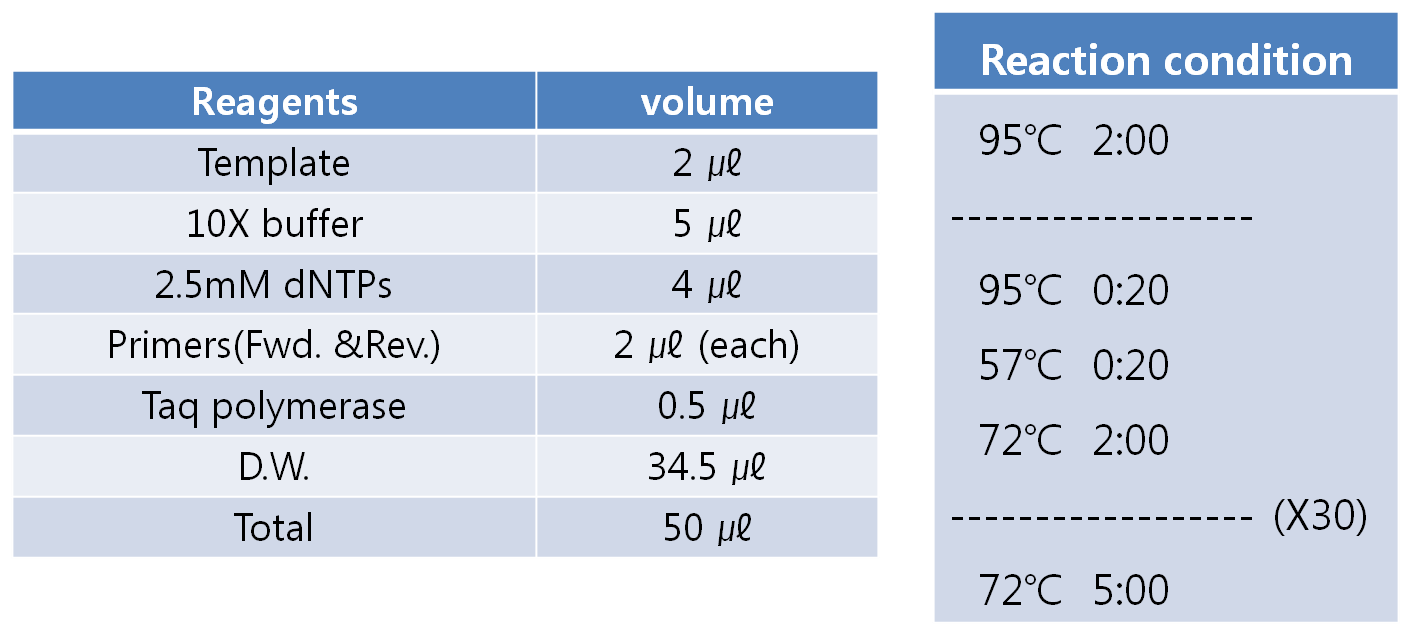
| - | ** Detail conditions are shown below.[[Image:PCR condition.jpg|center|thumb|400px]] | + | *** Detail conditions are shown below.[[Image:PCR condition.jpg|center|thumb|400px]] |
| + | **Treatment of PCR products | ||
| + | ***Method 1. | ||
| + | # PCR products were gel-purified. | ||
| + | # Digested with DpnI for 2~3 hours. | ||
| + | # Repurified and suspended in elution buffer (10mM Tris, pH8.0) | ||
| + | ****Takes more time and product concentration is lower than method 2. But template vectors remain less. | ||
| + | ***Method 2. | ||
| + | # Add 1㎕ of DpnI to not purified PCR products. | ||
| + | # Incubate at 37℃ about 2~3 hours. | ||
| + | # This samples were gel-purified and suspended in elution buffer. | ||
| + | ****Takes less time and get high yield of products. Templates can be remained.(But it's not affect critically to transformation.) | ||
==== Notes ==== | ==== Notes ==== | ||
*Primers | *Primers | ||
Revision as of 08:00, 2 August 2010
Contents |
E. coli Gene Replacement
Lambda red recombinase method
(see this link - openwetware::gene replacement - for the method comparison)
Materials
- Plasmids
- pKD46 -sequence info.
- Coding lambda red recombinase
- Ampiciline resistance
- Temperature sensitive replication origin
- pKD13 - sequence info.
- 3.4kb, KanR flanking FRT sequence, R6K gamma replication origin
- Kanamycin resistance gene template plasmid
- pCP20
- Flipase system
- Ampiciline resistance
- Temperature sensitive replication origin
- pKD46 -sequence info.
- Reagents
- L-arabinose
- Lambda red recombinase pBAD promoter inducer
- L-arabinose
- Equipment
- Incubators (30°C and 37°C)
- Electroporator
Procedure
- Preparation of plasmids
- Grow up pKD46, pKD13, and pCP20 in proper host strains
- Perform minipreps to extract plasmids
- Transformation of pKD46 to target strain
- Preparation of competent cells
- Using the protocol of TSS method
- Preparation of competent cells
- Generation of cassette flanked by homologous arms
- Design of primers
- Primers have overhangs which were homologous to surrounding region of target gene.
- Sequences from cassette region have about 60℃ Tm-value.
- Overview is attached. File:Overview
- PCR condition
- Anneling temparature and extension time is depend on the target sequence.
- Anneling temparature is tested at 55℃ at first.
- Extension time is depend on the length of the target sequence. (1kb = about 1 min.)
- Detail conditions are shown below.
- Treatment of PCR products
- Method 1.
- Design of primers
- PCR products were gel-purified.
- Digested with DpnI for 2~3 hours.
- Repurified and suspended in elution buffer (10mM Tris, pH8.0)
- Takes more time and product concentration is lower than method 2. But template vectors remain less.
- Method 2.
- Add 1㎕ of DpnI to not purified PCR products.
- Incubate at 37℃ about 2~3 hours.
- This samples were gel-purified and suspended in elution buffer.
- Takes less time and get high yield of products. Templates can be remained.(But it's not affect critically to transformation.)
Notes
- Primers