Mesostate
From CSBLwiki
(Difference between revisions)
(→Mesostate) |
(→Mesostate) |
||
| Line 22: | Line 22: | ||
*Data production by [[배형섭]] | *Data production by [[배형섭]] | ||
*[http://roselab.jhu.edu/dist/manual/meso_lett.html Torsion angle mesostate] by LINUS (Rose Lab) | *[http://roselab.jhu.edu/dist/manual/meso_lett.html Torsion angle mesostate] by LINUS (Rose Lab) | ||
| - | http://www.pnas.org/content/102/45/16227/F1.large.jpg | + | [[file:F1.large.jpg|150px|thumb|Mesostate - it contains an error http://www.pnas.org/content/102/45/16227/F1.large.jpg]] |
| - | + | ||
====Calculation==== | ====Calculation==== | ||
Revision as of 08:54, 9 August 2010
|
Concept
- Protein structures (3D) can be parsed into a finite number of structural alphabets (e.g. a torsion angle of each residue)
- From these structural alphabets, we will identify meaningful structural words..
- Frequent structural words (motifs?) and popular sentences (super secondary structures?)
- Using these structural words, 1) building a profile, 2) relating evolution of molecules (proteins & folds) and organismic phylogeny (genomes; a catalog of words, sentences..)
- Assumptions - ...
- Expectation - Multiple Birth (Birth and death) model??
Procedure
Standard data set
- We are going to use the SCOP DB: sequences and structures in the Astral compendium
Torsion angles
- Structural alphabets based on torsion angle distributions in the ramachandran plot
images to be posted...
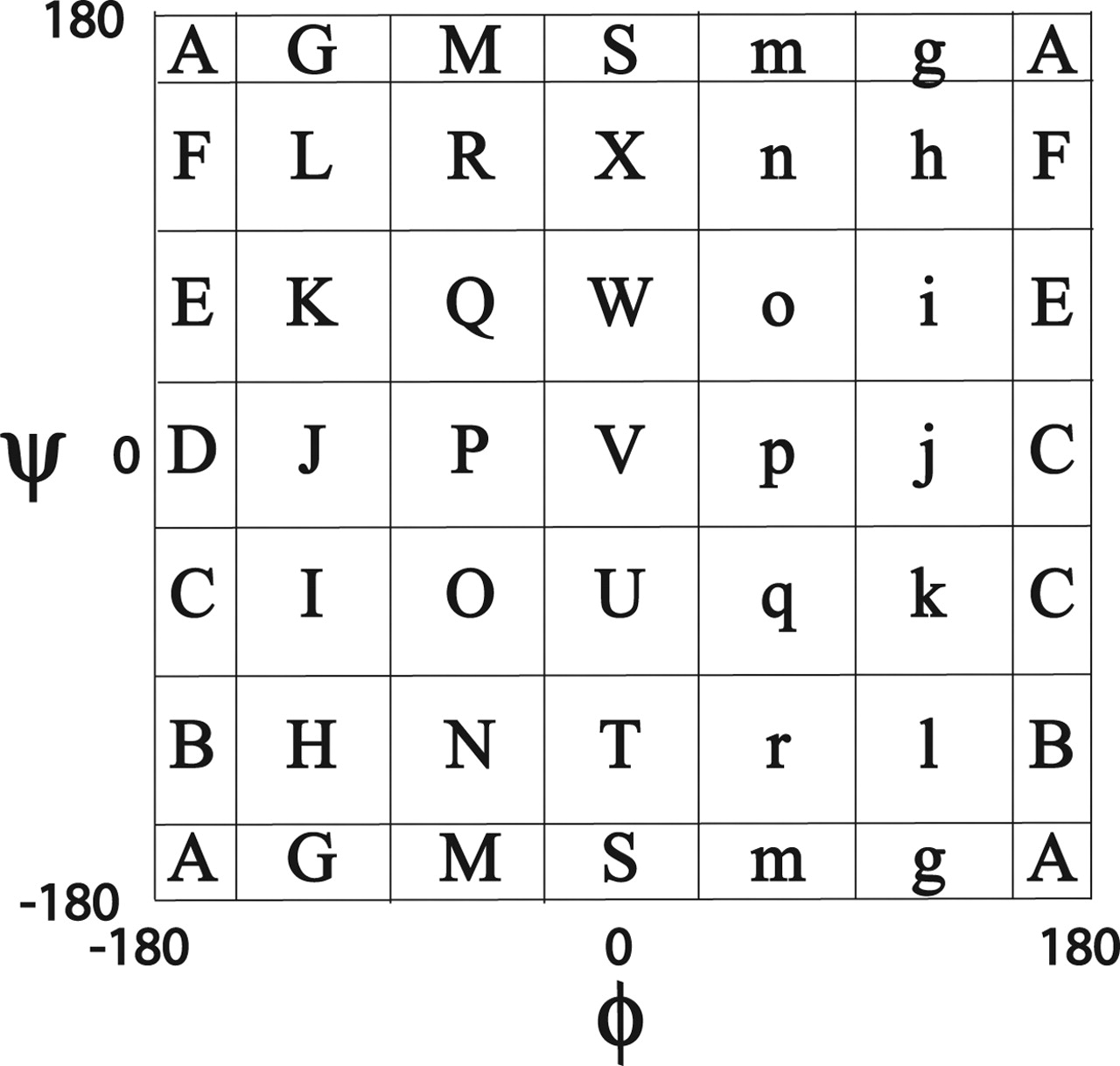
Mesostate
- Data production by 배형섭
- Torsion angle mesostate by LINUS (Rose Lab)
Calculation
- Following tools can be used to calculate torsion angles of backbones
Alphabet assignment
- Information theory
Profiling
- Normalization?
- Distance metric?
Applications
Status(result)
- TBA