Mesostate
From CSBLwiki
(Difference between revisions)
(→Status(result)) |
(→Status(result)) |
||
| Line 41: | Line 41: | ||
<pre> | <pre> | ||
# R script | # R script | ||
| + | <pre> | ||
| + | ## load saved R data | ||
| + | load("meso.rdata") | ||
| + | ## analysis | ||
| + | dim(meso) # dimension 11,810,116 residues | ||
| + | meso[1:2,] # check first two rows in the data (list) | ||
| + | dom = unique(meso$Domain) | ||
| + | ndom = length(dom) # 65,485 SCOP domains | ||
| + | nrow(meso)/ndom # average 180 residues (domain size) | ||
| + | # consider 1st, last residues are skipped.. | ||
| + | ## | ||
| + | ## ramachandran plot | ||
| + | ## | ||
| + | # randomly picking 5,000 residue's Phi & Psi | ||
| + | rn = sample(nrow(meso),5000) | ||
| + | png(file="ramachandran.plot") | ||
| + | plot(meso$Phi[rn],meso$Psi[rn],xlab="Phi",ylab="Psi",xlim=c(-180,180),ylim=c(-180,180),main="Ramachandran plot",col="gray") | ||
| + | # randomly picking 5,000 helices | ||
| + | rn = sample(which(meso$Structure=='H'),5000) | ||
| + | points(meso$Phi[rn],meso$Psi[rn],col="green") | ||
| + | # randomly picking 5,000 sheets | ||
| + | rn = sample(which(meso$Structure=='E'),5000) | ||
| + | points(meso$Phi[rn],meso$Psi[rn],col="red") | ||
| + | dev.off() | ||
| + | ## | ||
</pre> | </pre> | ||
*6 x 6 bins | *6 x 6 bins | ||
Revision as of 09:50, 29 August 2010
|
Concept
Procedure
Standard data set
- We are going to use the SCOP DB: sequences and structures in the Astral compendium
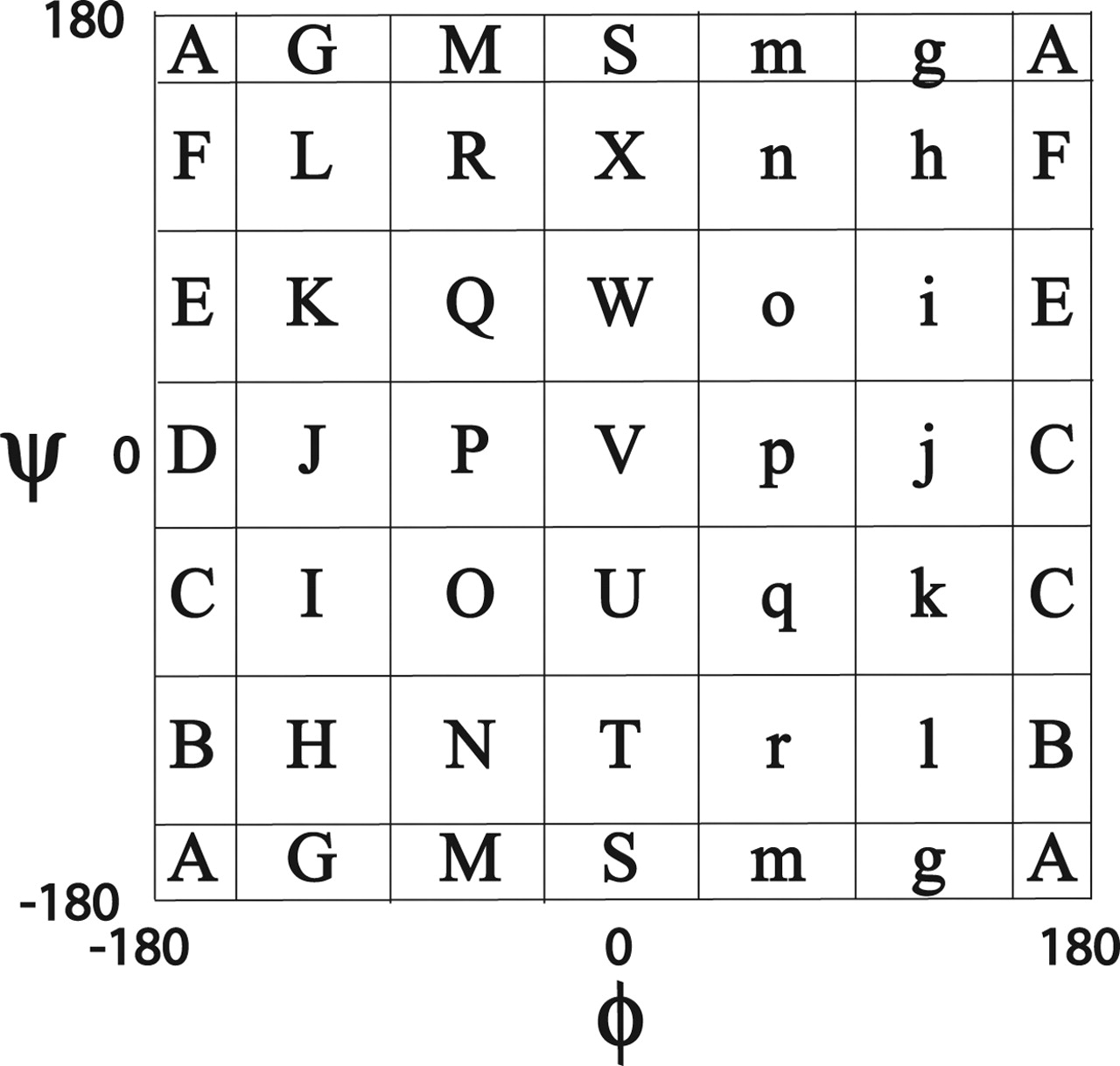
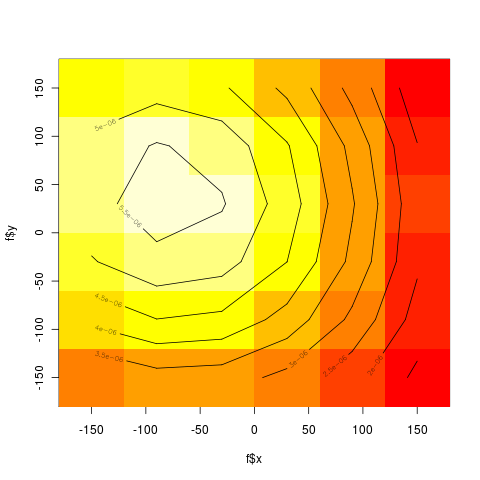
Torsion angles
- Structural alphabets based on torsion angle distributions in the ramachandran plot
images to be posted...
Mesostate
- Data production by 배형섭
- Torsion angle mesostate by LINUS (Rose Lab)
Calculation
- Following tools can be used to calculate torsion angles of backbones
Alphabet assignment
- Information theory
Profiling
- Normalization?
- Distance metric?
Applications
Status(result)
- Using R to check the data
# R script
<pre>
## load saved R data
load("meso.rdata")
## analysis
dim(meso) # dimension 11,810,116 residues
meso[1:2,] # check first two rows in the data (list)
dom = unique(meso$Domain)
ndom = length(dom) # 65,485 SCOP domains
nrow(meso)/ndom # average 180 residues (domain size)
# consider 1st, last residues are skipped..
##
## ramachandran plot
##
# randomly picking 5,000 residue's Phi & Psi
rn = sample(nrow(meso),5000)
png(file="ramachandran.plot")
plot(meso$Phi[rn],meso$Psi[rn],xlab="Phi",ylab="Psi",xlim=c(-180,180),ylim=c(-180,180),main="Ramachandran plot",col="gray")
# randomly picking 5,000 helices
rn = sample(which(meso$Structure=='H'),5000)
points(meso$Phi[rn],meso$Psi[rn],col="green")
# randomly picking 5,000 sheets
rn = sample(which(meso$Structure=='E'),5000)
points(meso$Phi[rn],meso$Psi[rn],col="red")
dev.off()
##
- 6 x 6 bins
> print(bins)
[,1] [,2] [,3] [,4] [,5] [,6]
[1,] 90545 18116 83067 120228 284531 1369188
[2,] 100305 84717 3685965 485872 653175 2066098
[3,] 4849 86683 1565006 5826 39358 294463
[4,] 23516 9348 2594 135671 25882 3456
[5,] 54491 12777 109935 234363 17244 41903
[6,] 35495 3524 13284 6641 5944 34984
References
Error fetching PMID 19188606:
- Error fetching PMID 19188606: