Molecular biology
From CSBLwiki
(Difference between revisions)
(→=Gene replacement) |
|||
| (3 intermediate revisions not shown) | |||
| Line 1: | Line 1: | ||
| - | {| align= | + | {| align=right cellpadding=15 |
|__TOC__ | |__TOC__ | ||
|} | |} | ||
==Reagents & Protocols== | ==Reagents & Protocols== | ||
| - | === | + | ===Recipes=== |
| - | *[[Chemicals]] | + | ;*[[Chemicals]] |
| - | *[[media:mba.pdf|Commonly used reagents and equipment in molecular biology (PDF)]] | + | ;*[[media:mba.pdf|Commonly used reagents and equipment in molecular biology (PDF)]] |
| - | *[[Concentration of antibiotics commonly used]] | + | ;*[[Concentration of antibiotics commonly used]] |
| - | *For DNA extraction (to remove some protein contaminants & inactivate enzymes): Phenol:Chloroform=1:1(v:v) solution | + | ;*For DNA extraction (to remove some protein contaminants & inactivate enzymes): Phenol:Chloroform=1:1(v:v) solution |
| - | *DNA sequencing check list | + | ;*DNA sequencing check list |
| - | + | :#[http://en.wikipedia.org/wiki/Phred_quality_score quality score] | |
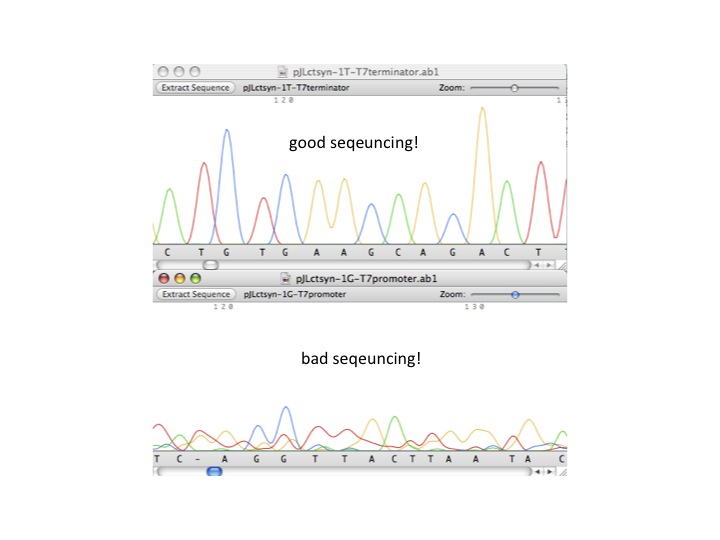
| - | + | :#good sequencing result vs. bad sequencing result (see below example) | |
| - | [[image:sequencingresult.jpg| | + | ::[[image:sequencingresult.jpg|center|thumb|good vs. bad sequencing example]] |
===Protocols=== | ===Protocols=== | ||
| Line 45: | Line 45: | ||
#Biological sequence editor [http://www.mbio.ncsu.edu/BioEdit/bioedit.html Bioedit] | #Biological sequence editor [http://www.mbio.ncsu.edu/BioEdit/bioedit.html Bioedit] | ||
#[http://en.wikipedia.org/wiki/Jmol Jmol] | #[http://en.wikipedia.org/wiki/Jmol Jmol] | ||
| + | ==Links== | ||
| + | *[http://www.ncbe.reading.ac.uk/ncbe/protocols/ NCBE practical protocols]: very good illustration for undergraduate students | ||
Latest revision as of 07:30, 16 January 2013
|
Reagents & Protocols
Recipes
- Chemicals
- Commonly used reagents and equipment in molecular biology (PDF)
- Concentration of antibiotics commonly used
- For DNA extraction (to remove some protein contaminants & inactivate enzymes): Phenol:Chloroform=1:1(v:v) solution
- DNA sequencing check list
- quality score
- good sequencing result vs. bad sequencing result (see below example)
Protocols
- PCR - General rules for primer design
- Genomic DNA extraction
- Competent cell preparation BSGC protocol (PDF)
- Easy & rapid competent cell preparation reference link - by HJKo
- Electrocompetent cell Protocol (PDF)
LIC cloning
- Reference:
Error fetching PMID 2235490:
- Error fetching PMID 2235490:
- LIC cloning introduction LIC introduction - Ligation Independent Cloning
- LIC cloning protocol (Berkeley Structural Genomics Center)
- Vector preparation
- Insert preparation
- LIC transformation:
- mix insert and vector in appropriate amounts
- incubate 5~30 min(20 min) at room temperature
- do transformation in E. coli (recommend to use electro-competent cells)
- LIC cloning protocol (Berkeley Structural Genomics Center)
Conjugation
- Donor: E. coli S17-1
Gene replacement
- E. coli knockout by the Wanner method (Lambda Red Recombinase)
Tools
webtools
softwares
- NIH Image: Gel analysis software
- Serial Cloner: DNA sequence management (PCR, restriction maps, etc.)
- BioX (Mac OS X only)
- Biological sequence editor Bioedit
- Jmol
Links
- NCBE practical protocols: very good illustration for undergraduate students